Introduction
The Microscopy Image Analyzer is a specialized GUI application I developed to address a common challenge in cell biology research: processing and presenting time-series microscopy data. When preparing microscopy images for scientific publications, researchers often need to track specific cells across multiple timeframes while maintaining consistent framing and presentation quality.
This tool streamlines the entire workflow from raw microscopy data to publication-ready figures and animations, eliminating the need for multiple software tools and manual processing steps.
Key Features
Image Sequence Handling
- Support for multiple image formats (JPEG, PNG, TIFF, BMP, GIF)
- Intuitive navigation through timeframe sequences
- Automatic sequence organization and management
- Built-in image format conversion capabilities
ROI Management
- Interactive ROI drawing with real-time preview
- Precise cell location targeting with coordinate input
- ROI resize and repositioning capabilities
- Batch processing across entire image sequences
Export Capabilities
- High-quality individual frame export
- Professional GIF animation creation
- MP4 video export with quality settings
- Smart file naming and duplicate handling
Application Interface
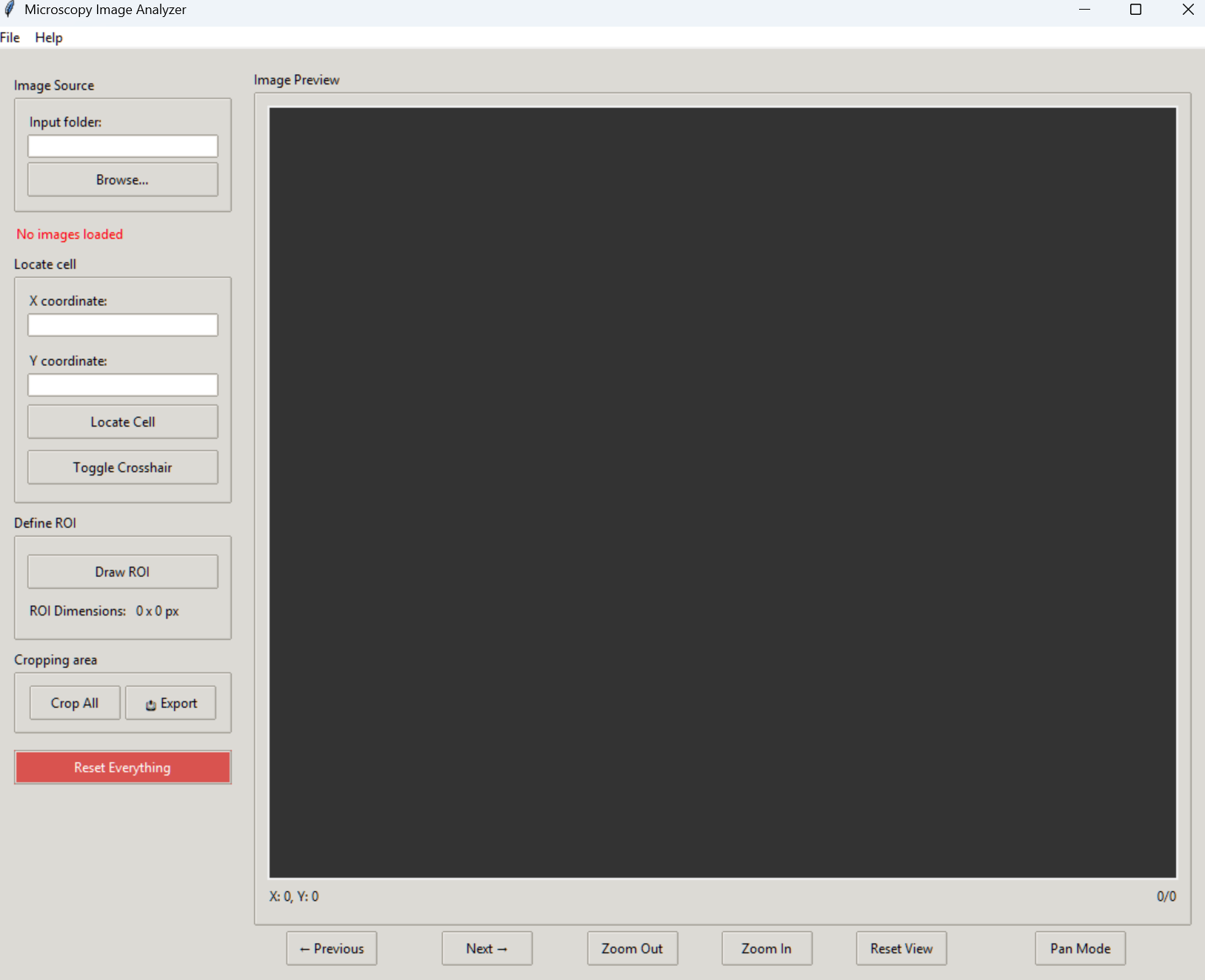
The main interface showing the image viewer, ROI tools, and control panels
Technical Implementation
Architecture Overview
The application follows a modular architecture with clear separation of concerns:
microscopy_analyzer/
├── src/
│ ├── core/ # Core processing logic
│ ├── gui/ # User interface components
│ └── utils/ # Helper functions
├── config/ # Configuration settings
└── tests/ # Unit testsGUI Components
The interface is built using tkinter with custom components for:
- Interactive canvas for image display and ROI manipulation
- Control panels for navigation and settings
- Professional export dialog with advanced options
- Status information display
Image Processing
The application leverages several Python libraries for image processing:
from PIL import Image, ImageTk
import numpy as np
import imageio
class ImageProcessor:
def __init__(self):
self.supported_formats = ['.jpg', '.jpeg', '.png', '.tiff', '.bmp', '.gif']
def load_sequence(self, folder_path):
"""Load and validate image sequence from folder"""
image_files = self._get_image_files(folder_path)
return [Image.open(f) for f in image_files]
def crop_with_roi(self, image, roi):
"""Crop image using ROI coordinates"""
x1, y1, x2, y2 = roi
return image.crop((x1, y1, x2, y2))
def create_animation(self, images, fps=5, output_path=None):
"""Create GIF animation from image sequence"""
if output_path is None:
output_path = 'animation.gif'
imageio.mimsave(output_path, images, fps=fps)Usage Guide
Installation
The application requires Python 3.7 or higher and can be installed using pip:
# Install required packages
pip install -r requirements.txt
# Run the application
python main.pyBasic Workflow
- Load Images: Select a folder containing timeframe images
- Navigate Sequence: Use Previous/Next buttons to browse images
- Define ROI: Draw region of interest around target cells
- Process Batch: Apply ROI to all images in sequence
- Export Results: Save cropped images or create animations
💡 Pro Tips
- Use coordinate input for precise cell targeting
- Preview animations before final export
- Utilize keyboard shortcuts for efficient navigation
- Save commonly used ROI coordinates for reuse
Applications
This tool is particularly valuable for:
📝 Manuscript Preparation
- Create consistently framed cell images
- Generate publication-quality figures
- Prepare supplementary animations
🔬 Cell Tracking
- Track cell movement over time
- Document morphological changes
- Create focused time-lapse sequences
🎓 Research Presentations
- Create engaging visual aids
- Demonstrate cellular processes
- Export presentation-ready media
Conclusion
The Microscopy Image Analyzer represents a significant improvement in microscopy image processing workflow, offering:
🎯 Key Benefits:
- Time Efficiency: Streamlined workflow reduces processing time
- Consistency: Maintains uniform presentation across image sequences
- Quality: Professional-grade output for publications
- Usability: Intuitive interface for researchers
🔮 Future Development
Planned enhancements include:
- Integration with machine learning for automated cell detection
- Advanced image analysis capabilities
- Support for additional file formats
- Enhanced batch processing features
